AUTOMATION WITH INTELLIGENCE

Introduction
In the past decade, we have seen development and advancement in using technologies in clinical research and life science. Some of key concepts are discussed in many articles and blogs and are evolving from database to dataverse, from Real-world data (RWD) to Real-world evidence (RWE), from digital simulation to digital twins, from metadata to metaverse, and from digital improvement, digital transformation to automation with intelligence (intelligence upgrade). All of these development and evolution are the results of standard development and and standard adoption, particularly in the life science.
The adoption of standards in the life sciences industry is accelerating, and with it, pharmaceutical and service companies are demanding greater efficiency and higher quality from standard-based solutions. Automation that is based on artificial intelligence (AI) and machine learning (ML) will be the next game changer in the industry to provide higher quality data and more efficient processes.
What are the challenges in managing standards adoption? How to identify potential approaches for automation through utilizing robotic processes and artificial intelligence? What kind of model can we use to assess maturity of adopting standards, improving efficiency in process and building intelligent automation in drug research?
Key Considerations for Automation
There are three main consideration for automation: standard adoption, code reusability and process repeatability.
Standard Adoption: Standard-based Integration
Delivering quality data is the core goal of clinical trials and data management in drug research. The key for data quality is to adopt a common standard, to adhere to the standard practices and processes, and to use industry-strength technology to reduce human errors. Adoption of standards not only increases interoperability and efficiency but also provides a foundation for data integration and increases the degree of code reusability.
The life-sciences industry has adopted Clinical Data Interchange Standards Consortium (CDISC) standards for clinical trial data and FDA electronic common technical document (eCTD) standards for document preparation for many years but still faces substantial challenges. FDA binding guidance went into effect on December 17, 2016, and requested sponsors whose studies start after December 17, 2016 must submit data in FDA-supported formats listed in the FDA Data Standards Catalog. The solutions based on these standards enable integration among siloed systems but integration solutions also need to be based on business requirements and provide end-to-end (E2E) intelligence for the business. Since some of the standards are still in the process of being developed or being matured, the changes about the standards – the metadata of the standards needs to be collected and managed properly and timely. So the more that standards are adopted, the more that meaningful and timely metadata are needed to manage the process of upgrading those standards and the application of the new standards to the existing processes and documentation.
Most of the data that are generated in clinical trials are stored and processed in a heterogeneous array of independent, non-interoperable systems. System integration is an engineering concept, the goal of which is to bring together the component subsystems into one system to deliver overarching functionality and to ensure that the subsystems function together as a single system. In the real world, system integration involves integrating existing, often disparate systems to increase value. Because Value = Quality / Cost, Value can be increased either by enhancing product Quality and hereby performance or by reducing the Cost to the customer.
As the number of independent systems increases, the number of integrations increases exponentially, whereas with a standard in place, the number of integrations increases linearly. This makes a compelling case for the use of standards. Data integration is a key element of conducting scientific investigations with modern platform technologies, and it is a starting point for the management of increasing complexity of drug discovery and clinical research, and for the eventual, full realization of economies of scale in larger enterprises that relies on reusable code that comes from data integration.
Code Reusability: Metadata-driven Automation
The key to reusability and automation is metadata. Metadata is data that describes other data and that provides information about other data. Metadata can be categorized into three main types: Descriptive metadata, Structural metadata, and Administrative metadata. The role of metadata for Life Sciences & Healthcare has changed: It is not only increasing the desire for integration across patient-centric business processes but also driving automation and collaboration within drug research among sectors in the life science industry.
There are two basic approaches to building reusable code: either opportunistic or planned. An opportunistic approach is an ad hoc way of finding or creating reusable code. A planned approach offers the opportunity to create reusable code in a systematic way that is the most efficient way possible. The code developed for datasets in one phase of a clinical study could be used for all phases or from one study that could be used for all studies in the same therapeutic area. It is a strategy for increasing productivity and improving quality in data transformation and standardization. Although simple in concept, successful code reuse implementation is difficult in practice.
There are many ways that we can increase the code reusability of an ETL process in the data integration. However, viewing data integration simply as a data issue or a technical issue underestimates the unique, scientific, management challenges that it embodies: challenges that could require significant process reengineering, methodological and even cultural changes in our approach to data, particularly metadata.
Process Repeatability: Improved Efficiency
Another important aspect of metadata-driven automation is process repeatability. Efficiency can be gained through building repeatable workflows. A defined and repeatable workflow also ensures work quality. However, in reality, there are many isolated systems and single purpose codes in the real world. How could reusable, silo codes be linked together into a repeatable process?
Important considerations for increasing code reusability include the following: 1) Adoption of standards is the key for code reusability, which involves the following aspects: Train people to understand the standards; Define standard templates; Build public libraries for code snippets and public transformation; Group code snippets and functional transformation into modular mapping and transformation; Define workflow to govern the process. 2) Metadata-driven processes are the key to automation because metadata is machine readable and makes the individual data sets meaningful by connecting them to provide new insights. 3) Automation helps to facilitate creation of utilities that replicate processes for Project Set Up, Mapping Specification, and Mapping Creation.
Once reusable components have been identified, we can follow the following steps to build reusable modules or even a base project which can be used to start a new project. Extract Common Components by 1) building a public code library that includes transformation and utilities (functions, procedures, packages, pluggable maps and workflows), 2) building a metadata repository that includes SDTM data modeling, controlled terminologies and specification look-up tables for mapping intelligence; and 3) creating a base project to build common modules and public locations (database links). Build Follow-on Projects: by creating location links to metadata repositories, importing public utilities through transformation and use of data rules and subject matter experts, and copying the base project and associated modules.
Intelligent Automation Models
From the discussion above, we can see that automation are from adoption of standards, metadata collection about the clinical study, and the process of conducting study; metadata-driven automation not only improves data quality but also increases efficiency. Collecting and classifying study-related metadata are the first steps to building artificial intelligence about clinical study automation.
Maturity Assessment Questions
| Questions | Indicator Level 1 | Indicator Level 2 | Indicator Level 3 | |
|---|---|---|---|---|
| Standards adoption (SA) | Is there at least one dedicated resource for standards adoption in the company? Is there periodical training for people involved in data processing and analyses? Is there a permanent governance body for standards adoption and implementation? Is there a metadata registration system implemented to manage standards adoption? Is there any robotic process automation (RPA) and artificial intelligence (AI) technology being used to gain insights and help with decisions? | 2 Yeses | 3~4 Yeses | 5 Yeses |
| Code reusability (CR) | Is there a common function library? Could the codes be modularized? Are code metadata being collected? | 1 Yes with 30% CR | 2 Yeses with 30~60% CR | 3 Yeses with above 60% CR |
| Process repeatability (PR) | Is there a defined workflow? Could workflow be grouped and classified? Is the process metadata being collected? | 1 Yes with 30% PR | 2 Yeses with 30~60% PR | 3 Yeses with above 60% PR |
Conceptual Model
Intelligent automation is a more advanced form of what is commonly known as robotic process automation (RPA) with contextual metadata. The RPA is driven by predefined contextual metadata such as how to log into various systems, when to conduct pivot transformation, how to merge data from different domains, etc. This type of operation may be overwhelming to end users but machines have different strengths and capabilities that complement their human supervisors. Together, they’re changing what’s possible.
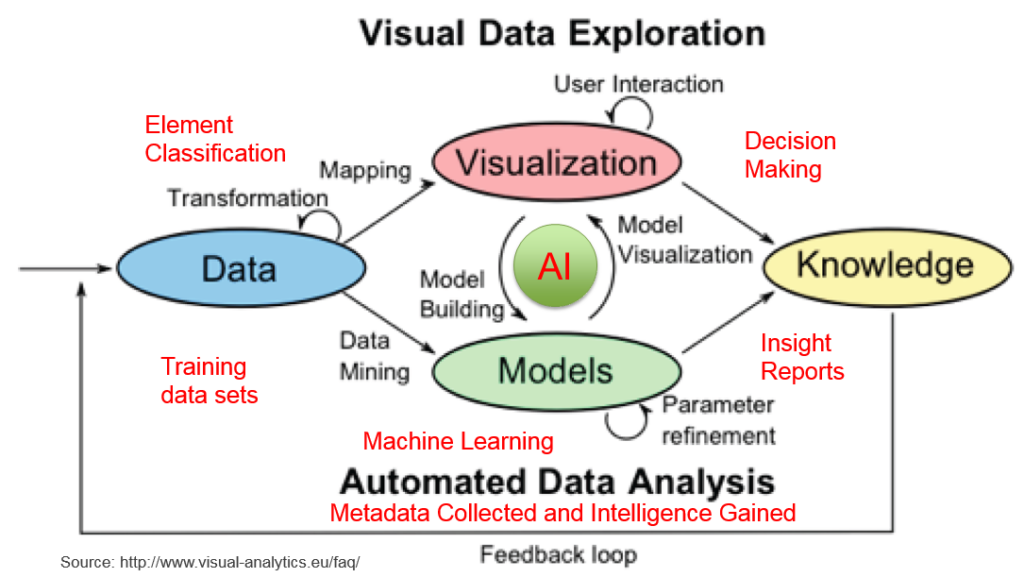
The conceptual model depicts how data is processed, classified, analyzed, modeled, visualized and used in gaining new insightful knowledge and making intelligent decisions. It has four components:
- 1) Data Processing: to make sure the data are clean, classified and ready to be used;
- 2) Machine learning: to build meaningful algorithms or models such as decision tree, deep neural network, convolutional neural network, recurrent neural network, etc.
- 3) Data Presenting: to present data in different ways and different dimensions to allow users to understand the meanings and insights of the data and to help making decisions. Kelly Lautt at Microsoft in 2007 created a term – data presentation architecture (DPA) and recognized that it requires a much broader skill set than data visualization. Data visualization is just one element of DPA ;
- 4) Intelligence Building: to feed more diverse data, collect more metadata, gain more insights and make better decisions.
Maturity Model
A maturity model of intelligent automation is proposed to evaluate a company’s maturity level with three suggested key indicators:
- 1) Standards adoption (SA): this could be the potential integration indicator.
- 2) Code reusability (CR): is to make clinical research more efficient and can be used as an efficiency indicator.
- 3) Process repeatability (PR): is to make conducting clinical research more intelligent and can be used as an intelligence indicator.
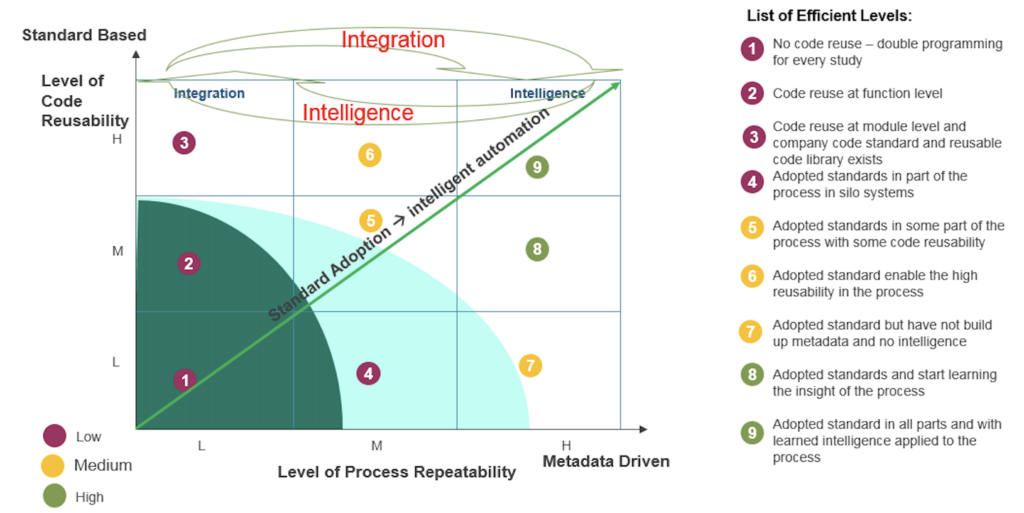
Standard-based systems allow for integration while metadata-driven systems enable automation; As more intelligence is collected about clinical data, a higher level of integration is achieved.
Further automation can be achieved through “Intelligent Data Flow”. Code reusability and process repeatability make data conversion very fast; the compliance check ensures the quality is good; thus, Intelligent Data Flow makes the whole clinical data lifecycle smart! So it makes the overall project relatively cheap.
Maturity Descriptive Model and Levels
Based on the answers to the questions proposed, each key indicator can be divided into three levels. These three indicators can be used as X (PR), Y (CR), and Z (SA) axles to form a cube to show a matrix for efficiency, based on metadata-driven automation – level of process repeatability (X), standard-based integration – level of code reusability (Y), and adoption of standards – level of intelligent automation (Z). It shows that standard-based systems allow for higher levels of integration measured by code reusability while metadata-driven systems enable higher levels of automation measured by process repeatability. The more that standards are adopted, the more that meaningful metadata could be applied in the process and the more efficient the process could become.
It may not be possible to use quantitative measurements to clearly define each maturity level (ML), but with the three levels in each key indicator, automation efficiency can be divided into 9 maturity levels (MLs) based on the combination of standards adoption, code reusability and process repeatability in the table – definition of indicator levels in the mutuality model.
- ML1: No common standard is adopted; No codes are reused, double programming for every study; It is difficult to reproduce the process in another study.
- ML2: Departmental standards exist in the company; Small sets of functions being developed and codes are reused at functional level; Processes used in different studies are evaluated but process repeatability is not addressed.
- ML3: Coding standard and styles exist but no common data and process standard being adopted; Common functions and macros are developed, code reuse at module level and companywide code standard and reusable code library exist; Process repeatability is fragmented and very manual.
- ML4: Begin standards education and adopt standards in part of process in siloed systems; Still conduct double programming for every study with small percentage of codes being reused; Data flow in some phases of the study have been defined and about half of the process can be repeated cross studies in the same therapeutic areas.
- ML5: A data standard coordinator is pointed and data standard is adopted in some parts of the whole study life cycle; Some reusable functions and modules are built and put into code libraries and are used cross studies; Some common workflows are defined and used cross studies.
- ML6: Dedicated standard personnel has been hired to manage standards adoption; Most functions conducting data transformation, standardization and classification are put into a shared common library and can be used for most of the studies; Some common workflows are defined and used cross studies in multiple therapeutic areas.
- ML7: Standard governance body is formed and standard department is established to coordinate all the standard implementation in the company; Double programming for every study is conducted with some code reusability; some algorithmic metadata have been collected but no metadata registration system (MDR) is built; Collected metadata help to automate the most of the workflows being defined and used cross studies.
- ML8: Standard governance body is formed and standard department is established to coordinate all the standard implementation in the company; a MDR system is implemented; Some reusable functions and modules are built and put into code library and are used cross studies; some visualization tools are used to explore the data; Collected metadata help to automate the most of the workflows being defined and used cross studies.
- ML9: Standards adoption is championed by the senior executives in the company; the governance body is embedded in all parts of the process; training on standards adoption is conducted periodically; a metadata repository system is implemented to manage all versions of the standards, code libraries, workflow definitions and their usage in various studies. 100% code reusability achieved through building modules, models, and services. 100% process repeatability achieved through collecting metadata, using metadata to drive the whole process.
Smart Automation in Bio Analytics
A key element of application-centric artificial intelligence is context with three main areas: smart classification, smart recognition, and smart predictions that encompass many cutting-edge AI and machine learning capabilities.
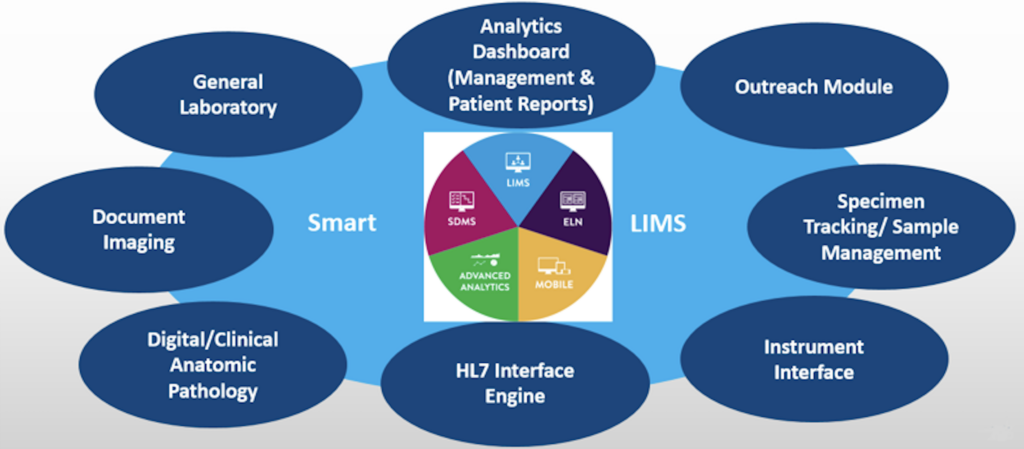
Many contract research organizations (CROs) have turned to technologies to build a smart lab with various systems designated to acquire, process and analyze lab data. Features of smart labs are shown in Figure above. Commonly deployed systems include:
- 1) Laboratory Information System (LIS) for processing and reporting data related to individual patients in a clinical setting;
- 2) Laboratory Information Management System (LIMS) for process and report data related to batches of samples from biology labs, water treatment facilities, drug trials, and other entities that handle complex batches of data.
- 3) Electronic Laboratory Notebook (ELN): for documenting lab research work with electronic notes that are searchable, shareable and have safeguards of security and backup.
- 4) Lab Execution System (LES): for quality control, quality assurance and compliance in laboratories.
- 5) Scientific Data Management System (SDMS): for acting as a document management system (DMS), capturing, cataloging, and archiving data generated by laboratory instruments (HPLC, mass spectrometry) and applications (LIMS, ELN). It is a gatekeeper, serving platform- independent data to informatics applications and/or other consumers.
These systems are designed to gather, store, and analyze large volumes of data, and some drug discovery units in particular have taken the lead in automating and robotizing their research laboratories. It may take a long time for a fully automated lab to be developed but it is possible to have micro robots in a chip to do very complicated experiments and analysis using microelectromechanical system [MEMS] techniques. There are in place smart kiosks or box configurations, such as bioanalytical wet chemistry kiosk (BWCK), bioanalytical wet chemistry in a box (BWCIB), and the bioanalytical laboratory in a box.
In general, lab automation refers to the use of technology to streamline or substitute manual manipulation of equipment and processes. This field of lab work comprises different automated lab instruments, devices, software algorithms and methodologies used to enable, expedite and improve efficiency and enhance effectiveness of scientific research and test analysis. The bioanalytical wet-chemistry automation of existing robotic platforms can be described as SMART systems: Simple to use; Mindful of user error; Assay class automation; Robust, rugged and reliable; Tied into organization’s IT systems. These systems run in a validated environment and are compliant with 21 CFR Part 11 and good laboratory practice (GLP). As Bungers has pointed out, the smart lab of the future will be made of flexible, digital integration, automation and robotics, integrated functional surfaces and modular systems.